Georgia Tsiliki1 †, Cristian R. Munteanu2,3 †, Jose A. Seoane4, Carlos Fernandez-Lozano2, Haralambos Sarimveis1 and Egon L. Willighagen3
†Equal contributors, 1School of Chemical Engineering, National Technical University of Athens, Athens, Greece, 2Computer Science Faculty, University of A Coruña, Spain, 3Department of Bioinformatics-BiGCaT, NUTRIM, Maastricht University, Maastricht, The Netherlands, 4Stanford Cancer Institute, Stanford University, Palo Alto, CA, USA
ABSTRACT
Background
Predictive regression models can be created with many different modelling approaches. Choices need to be made for data set splitting, cross-validation methods, specific regression parameters and best model criteria, as they all affect the accuracy and efficiency of the produced predictive models, and therefore, raising model reproducibility and comparison issues. Cheminformatics and bioinformatics are extensively using predictive modelling and exhibit a need for standardization of these methodologies in order to assist model selection and speed up the process of predictive model development. A tool accessible to all users, irrespectively of their statistical knowledge, would be valuable if it tests several simple and complex regression models and validation schemes, produce unified reports, and offer the option to be integrated into more extensive studies. Additionally, such methodology should be implemented as a free programming package, in order to be continuously adapted and redistributed by others.
Results
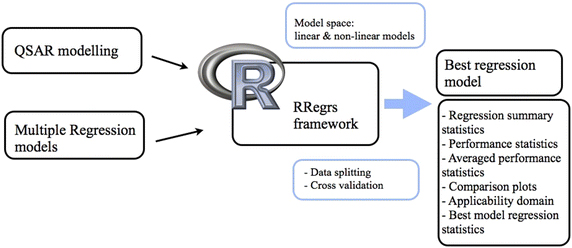
We propose an integrated framework for creating multiple regression models, called RRegrs. The tool offers the option of ten simple and complex regression methods combined with repeated 10-fold and leave-one-out cross-validation. Methods include Multiple Linear regression, Generalized Linear Model with Stepwise Feature Selection, Partial Least Squares regression, Lasso regression, and Support Vector Machines Recursive Feature Elimination. The new framework is an automated fully validated procedure which produces standardized reports to quickly oversee the impact of choices in modelling algorithms and assess the model and cross-validation results. The methodology was implemented as an open source R package, available at https://www.github.com/enanomapper/RRegrs, by reusing and extending on the caret package.
Conclusion
The universality of the new methodology is demonstrated using five standard data sets from different scientific fields. Its efficiency in cheminformatics and QSAR modelling is shown with three use cases: proteomics data for surface-modified gold nanoparticles, nano-metal oxides descriptor data, and molecular descriptors for acute aquatic toxicity data. The results show that for all data sets RRegrs reports models with equal or better performance for both training and test sets than those reported in the original publications. Its good performance as well as its adaptability in terms of parameter optimization could make RRegrs a popular framework to assist the initial exploration of predictive models, and with that, the design of more comprehensive in silico screening applications.
(for full presentation, please visit this link)
